Advantages
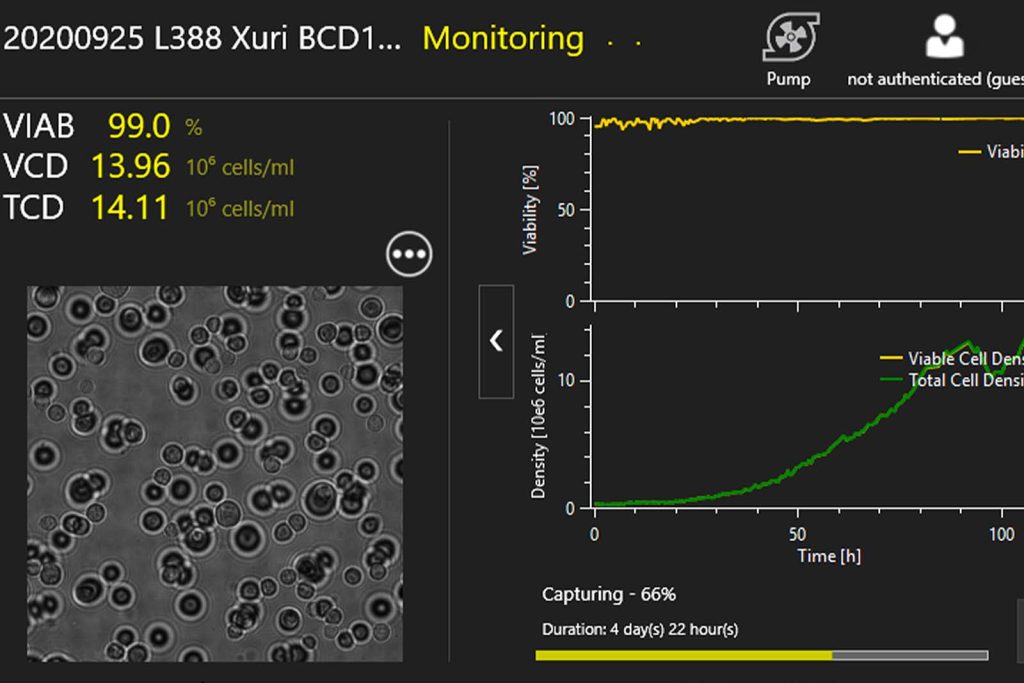
Real-time & automated
Closed-loop, online sampling allows for real-time process control
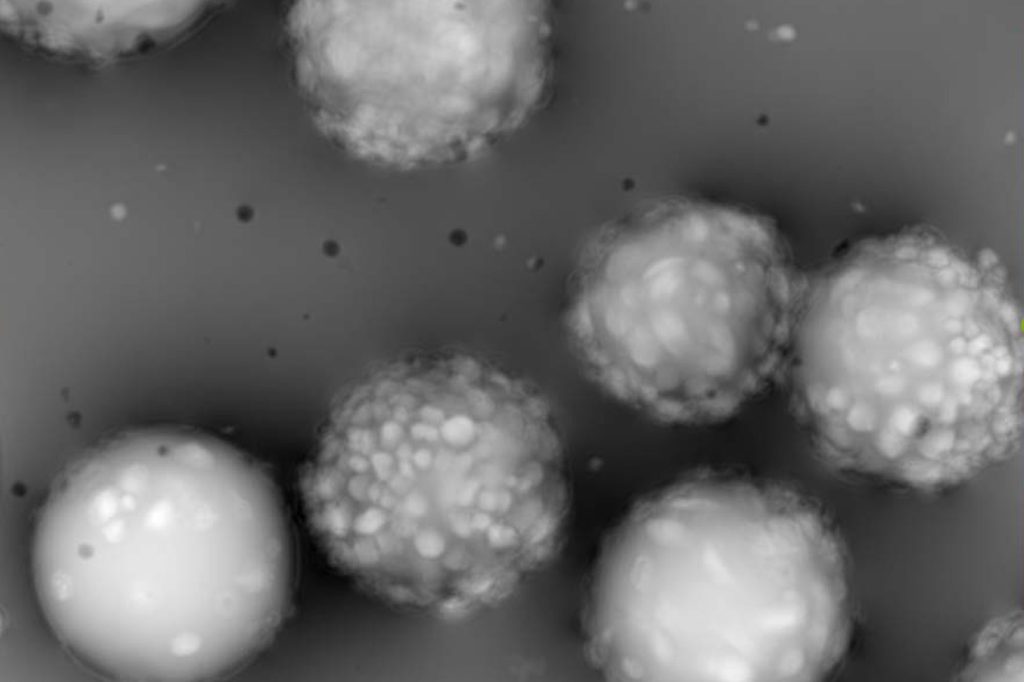
Label-free & non-destructive
No labeling is required; the cells are analyzed and return to the culture without damage

From development to manufacturing
Our solutions are adapted to every stage of bioprocess development, from small volumes to GMP manufacturing
A robust image-based process control & optimization platform
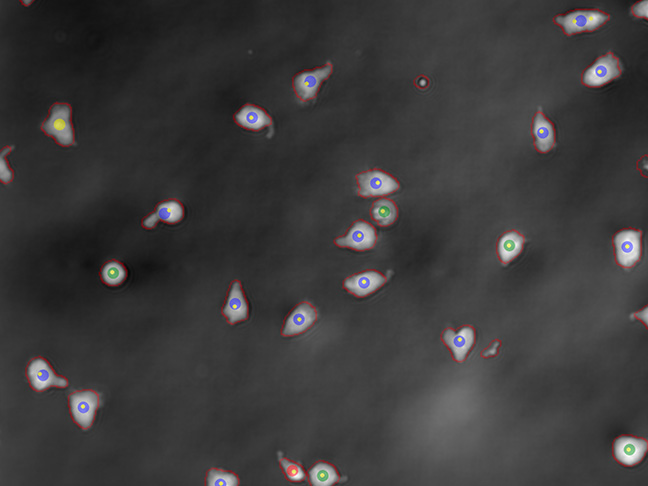
Activation
Real time online readout of the activation ratio of your cells in a non-destructive way, for process optimisation in PD and process decisions in GMP manufacturing.
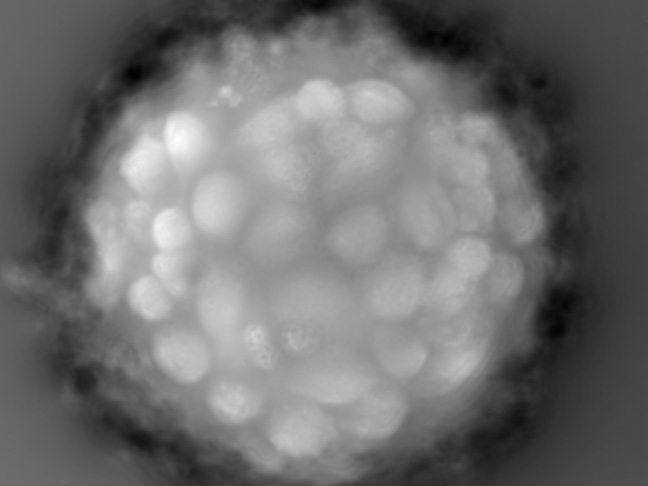
Microcarrier metrics
Real-time assessment of TCD and VCD (both on floating and attached cells) of cells grown on Dextran-based microcarriers, kinetics insight (attachment, detachment), as well as microcarriers quality control (diameter, defect detection.
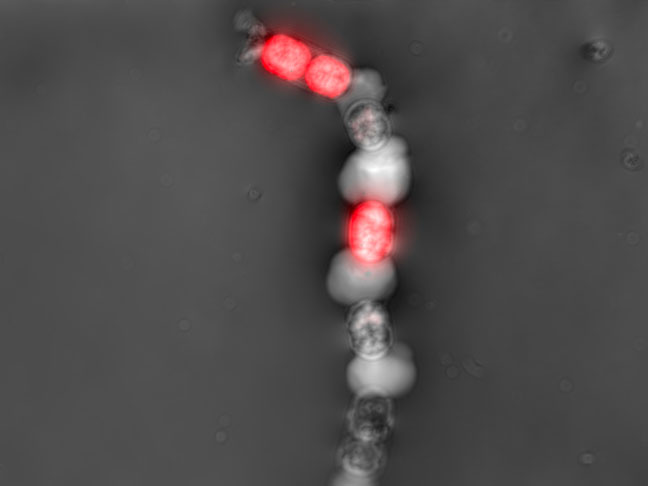
Contamination detection
Early detection of bacterial or fungal contamination of your culture while you monitor your preferred CPP
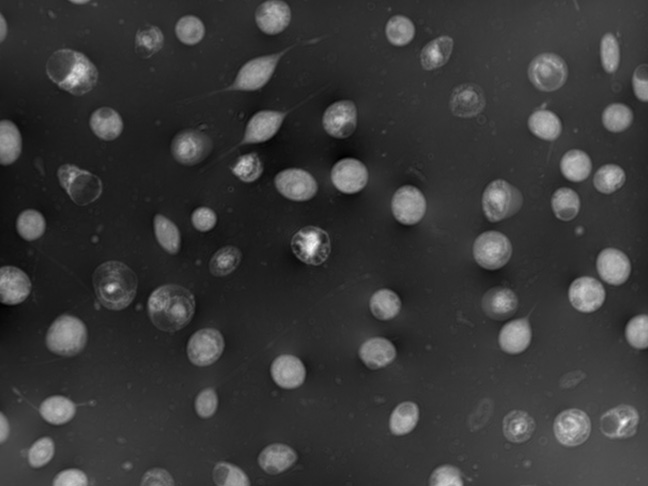
Viral infection kinetics
Keep a close watch on viral infection of SF9 and HEK cells and predict cell density, ensuring your gene-therapy process is always on track.
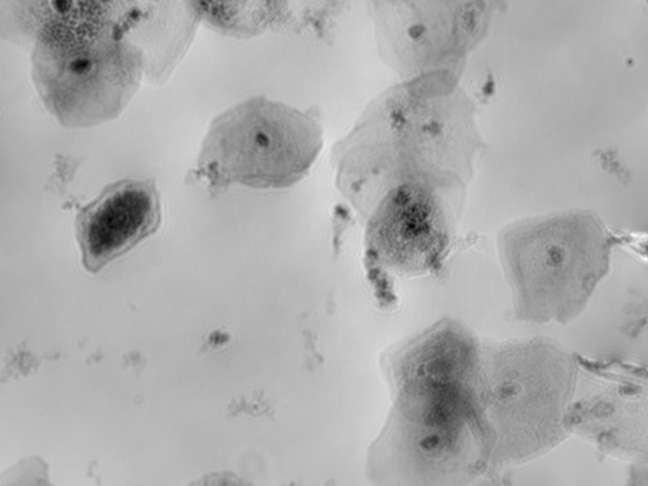
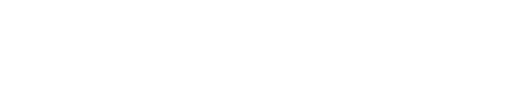
TCD & VCD
Automated online monitoring of TCD & VCD for CART, CAR-NK, SF9, HEK, and many other cell types.
Case studies
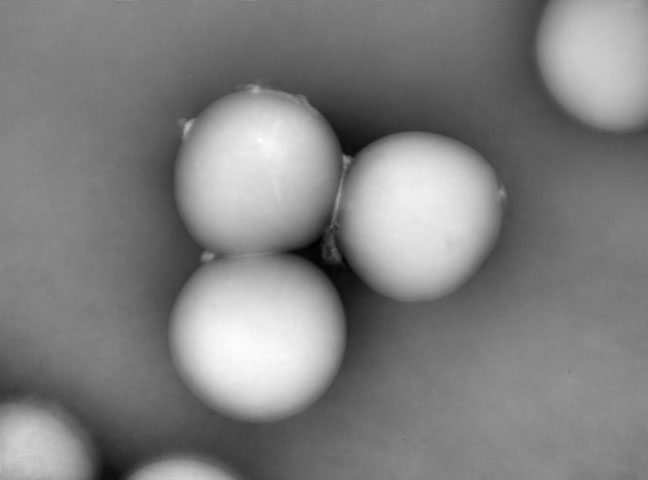
Automation of T-cell Expansion Using the iLine F.
A joint talk with Bristol Myers Squibb presented at the ASGCT digital event held on May 14, 2020.
Discover the case



